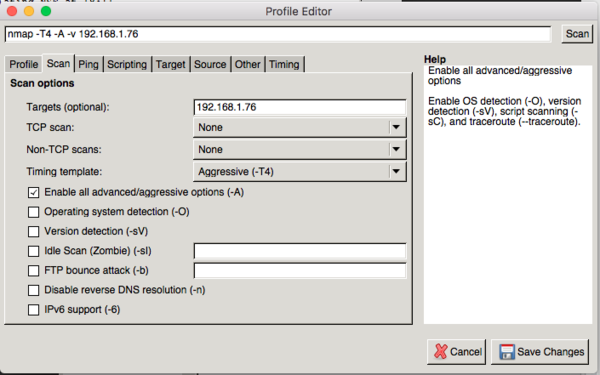
This is good to know if you are securing your server.
#Zenmap profiles Pc
#Zenmap profiles install
I will leave it up to you to install the packages. To run Zenmap, you need nmap installed first. You will be prompted for the root password. From the Linux command prompt type : " sudo zenmap". In order for Zenmap to fully operate, you must run it as root. It will then scan the server's ports and produce results based on what it found. Zenmap is run from a user's computer and pointed at a server's IP address or domain name. Nmap and Zenmap only work on Linux based systems such as CentOS, Redhat or Ubuntu. This webpage will demonstrate the basic capabilities of Zenmap. Command line is not fun but the good thing is that there is a free official GUI available called Zenmap. Nmap is a free Linux command line tool used for scanning a server's network connection to see which ports are exposed. The NMAP data will be deposited on the NCBI GEO database and our own SPIED web tool and should prove a useful resource for the wider research community.Zenmap Port Scanning for Security - TelecomWorld 101 Further, 30 of the NMAP profiles show anti-correlation with AD associated profiles.

The compound IPSC profiles showed a significant correlation with the corresponding CMAP profiles resulting in a significant positive enrichment in rank order when the IPSC profiles were queried against CMAP, demonstrating a degree of consistency across cell types. A set of 158 compounds with CMAP profiles tending to be the reverse of those associated with AD were assayed for their transcriptional effects on human IPSC derived cortical neuronal cultures. These profiles were shown to be highly consistent and viable queries for a CMAP repositioning approach. Various transcriptional profiles derived from post mortem AD samples and murine models of AD were generated based on publicly available data.
#Zenmap profiles full
To investigate the full neuronal transcriptional landscapes of a set of compounds targeted to Alzheimer's disease (AD) we have built NMAP (Neuronal Connectivity Map) a database of drug induced transcriptional profiles in human IPSC derived cortical neurons. The CMAP signatures, however, were generated in cancer cell lines and the LINCS data, defined over a wider set of cell types, is based on a limited set of 1000 landmark gene responses.

The Broad institute's Connectivity Map (CMAP) and Library of Integrated Network-Based Cellular Signatures (LINCS) projects have generated a large library of cellular transcriptional signatures in response to drug, genetic, and disease perturbation.
